What'sNEW July-September 2012
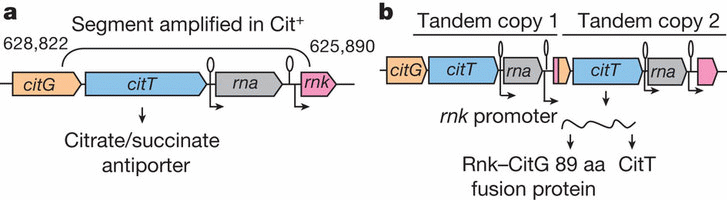
Richard Lenski's research group has analysed the evolution of aerobic citrate metabolism among cloned bacteria, a development first reported in 2008. It looks like an example of macroevolutionary progress in a quarantined system, so understanding how it happened is important. Now the same group reports that a previously silent genetic subroutine and an active regulatory sequence were duplicated and recombined into a configuration that became functional.
"The Cit+ trait originated in one clade by a tandem duplication that captured an aerobically expressed promoter for the expression of a previously silent citrate transporter." Also, point mutations before and after the duplication and recombination enabled and refined the new metabolic capability. These mutations were individually identified as part of the long and challenging study.
After reading the new analysis, our initial reaction is that the "cryptic gene" scenario, offered in 2008 as a possible explanation for the development, is the correct one — the "previously silent citrate transporter" being that gene. The recruited promoter, before getting its new assignent, was already active in a related role. The point mutations that enabled the first steps simply explored the genetic potential already present. The ones that refined the citrate metabolism exemplify optimization within a limited range. All of this would be consistent with both darwinism and cosmic ancestry. And none of it, in our opinion, demonstrates sustainable macroevolutionary progress in a quarantined system. We will stay tuned for additional news and informed comments.
 Zachary D. Blount, Jeffrey E. Barrick, Carla J. Davidson and Richard E. Lenski, "Genomic analysis of a key innovation in an experimental Escherichia coli population" [link], doi:10.1038/nature11514, p513-518 v489, Nature, 27 Sep (online 19 Sep) 2012. Zachary D. Blount, Jeffrey E. Barrick, Carla J. Davidson and Richard E. Lenski, "Genomic analysis of a key innovation in an experimental Escherichia coli population" [link], doi:10.1038/nature11514, p513-518 v489, Nature, 27 Sep (online 19 Sep) 2012.
 Evolutionary innovation caught in the act by Hristio Boytchev, The Washington Post, online 19 Sep 2012. Evolutionary innovation caught in the act by Hristio Boytchev, The Washington Post, online 19 Sep 2012.
 Bacteria's Key Innovation Helps Understand Evolution, The National Science Foundation (also EurekAlert!), 19 Sep 2012. Bacteria's Key Innovation Helps Understand Evolution, The National Science Foundation (also EurekAlert!), 19 Sep 2012.
 Richard Lenski, homepage at Michigan State University. Richard Lenski, homepage at Michigan State University.
 Cloned bacteria evolved an unexpected feature is our What'sNEW article about this evolutionary development, posted 5 Jun 2008. Cloned bacteria evolved an unexpected feature is our What'sNEW article about this evolutionary development, posted 5 Jun 2008.
 ...Is Evolutionary Progress in a Closed System Possible? is the related webpage with special What'sNEW section for Lenski et al. ...Is Evolutionary Progress in a Closed System Possible? is the related webpage with special What'sNEW section for Lenski et al.
 Testing Darwinism versus Cosmic Ancestry is a related local webpage. Test One: Biology mentions the experiment by Lenski et al. Testing Darwinism versus Cosmic Ancestry is a related local webpage. Test One: Biology mentions the experiment by Lenski et al.
 Thanks, Bob Sweeney, for a first alert. Thanks, Bob Sweeney, for a first alert.
 Michael Behe agrees with our assessment by email, 24 Sep 2012. Michael Behe agrees with our assessment by email, 24 Sep 2012.
 Rose-Colored Glasses: Lenski, Citrate, and BioLogos by Michael Behe, Evolution News, 13 Nov 2012. Rose-Colored Glasses: Lenski, Citrate, and BioLogos by Michael Behe, Evolution News, 13 Nov 2012.
Neil DeGrasse Tyson, Director of the Hayden Planetarium, endorses panspermia and discusses some of its implications.
 ...A fascinatingly disturbing thought, Dr. Neil DeGrasse Tyson in a 12.5 minute YouTube video, posted 25 Oct 2011 (also on Universe Today, posted 29 Mar 2013.) ...A fascinatingly disturbing thought, Dr. Neil DeGrasse Tyson in a 12.5 minute YouTube video, posted 25 Oct 2011 (also on Universe Today, posted 29 Mar 2013.)
 Thanks, twice, Google Alerts. Thanks, twice, Google Alerts.
 What Difference Does It Make? We suppose that cosmic ancestors might seem godlike. Tyson does, too. What Difference Does It Make? We suppose that cosmic ancestors might seem godlike. Tyson does, too.
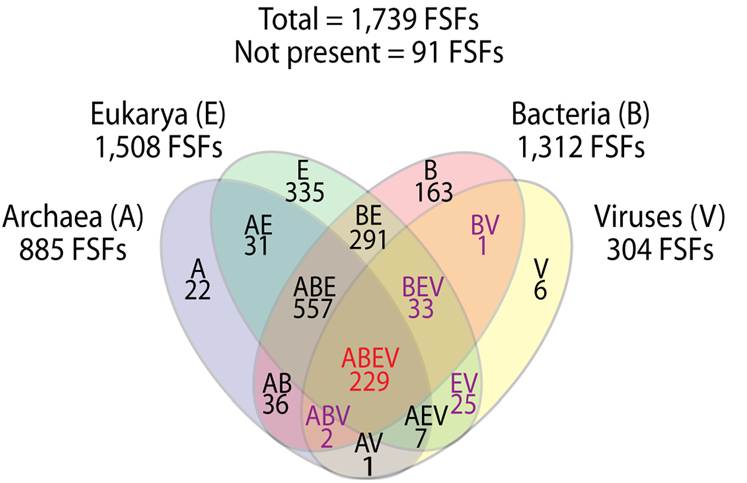
Very large viruses coexisted with or preceded the first primordial cells, according to three bioinformaticians affiliated with the University of Illinois, Urbana. The three also conclude that the large viruses were once even larger, but have lost many of their original genes. "Thus, mimiviruses (and megaviruses) should be viewed as the least reduced forms of an ancestral virus or cell that either coevolved or predated the cellular ancestors."
FSFs are not equally distributed in the proteomes of Archaea (A), Bacteria (B), Eukarya (E), and viruses (V). In turn, FSFs exist that are uniquely present (groups A, B, E or V) or are shared by two (AB, AE, AV, BE, BV, EV), three (ABE, ABV, AEV, BEV) or all (ABEV). |
Using new computer algorithms, the three compared related protein domains — fold families and fold super-families (FSFs) — among archea, bacteria, eukaryotes, and the large viruses. While many FSFs were unique to one or another category, the number shared between viruses and the others, especially unicellular parasites, was significant. The most parsimonious conclusion is that large viruses are at least as old as cells, and once contained even more genes than they do today. This would make viruses a fourth superkingdom of life.
These conclusions do not easily fit into standard darwinian theory, in which life on Earth began as a cell with a few genes and added more genes later. But the analysis readily suits and supports cosmic ancestry. Moreover, it suggests an even greater role for viruses than we formerly proposed.
 Arshan Nasir, Kyung Mo Kim and Gustavo Caetano-Anolles, "Giant viruses coexisted with the cellular ancestors and represent a distinct supergroup along with superkingdoms Archaea, Bacteria and Eukarya" [Open Access abstract], doi:10.1186/1471-2148-12-156, v12 n156, BMC Evolutionary Biology, 24 Aug 2012. Arshan Nasir, Kyung Mo Kim and Gustavo Caetano-Anolles, "Giant viruses coexisted with the cellular ancestors and represent a distinct supergroup along with superkingdoms Archaea, Bacteria and Eukarya" [Open Access abstract], doi:10.1186/1471-2148-12-156, v12 n156, BMC Evolutionary Biology, 24 Aug 2012.
 Viruses and Other Gene Transfer Mechanisms is a related local webpage. Viruses and Other Gene Transfer Mechanisms is a related local webpage.
What'sNEW about HGT  | |
 Metazoan Genes Older Than Metazoa? is a related local webpage. Metazoan Genes Older Than Metazoa? is a related local webpage.
 Genes Older Than Earth? is a related local webpage. Genes Older Than Earth? is a related local webpage.
 ...More functional protein domains than the first cells! is about work by members of the same team, What'sNEW, 26 May 2011. ...More functional protein domains than the first cells! is about work by members of the same team, What'sNEW, 26 May 2011.
 The protein was there to begin with.... describes related work by members of the same team, What'sNEW, 7 Oct 2011. The protein was there to begin with.... describes related work by members of the same team, What'sNEW, 7 Oct 2011.
 The standard RNA World theory is disputed... describes work using related methods, What'sNEW, 13 Mar 2012. The standard RNA World theory is disputed... describes work using related methods, What'sNEW, 13 Mar 2012.
...The ability of genes to function perfectly well across species boundaries has resulted in a significant horizontal flow of genes. Whether the genes are transferred by transposons, viruses, bacteria, or other vectors, or perhaps through direct contact or initial hybridization-like events, the horizontal flow of genes is a part of the story of life.
One of the major theoretical debates raised by HGT is the validity of the notion of the tree of life. ...One needs to be continually reminded that submitting multiple sequences ...to phylogenetic analysis produces trees because that is the nature of the algorithms used.
Homoplasy is traditionally defined as the sharing of a character between two species that [was] not possessed by their last common ancestor. ...Traditionally, it has been assumed that homoplasy is the result of convergent evolution, parallel evolution, or reversion to an ancestral state.... HGT can give rise to homoplasy as well....
If we accept that eukaryotes are chimeras built upon chimeras, we can expand the parameters within which to construct theories of eukaryotic origins.

These words come from a new paper by biochemist and microbiologist Michael Syvanen of UC Davis, an expert on horizontal gene transfer.
 Michael Syvanen, "Evolutionary Implications of Horizontal Gene Transfer" [abstract | 15 abstracts], doi:10.1146/annurev-genet-110711-155529, v 46 p341-358, Annual Reviews Genetics, Dec (online 29 Aug) 2012. Michael Syvanen, "Evolutionary Implications of Horizontal Gene Transfer" [abstract | 15 abstracts], doi:10.1146/annurev-genet-110711-155529, v 46 p341-358, Annual Reviews Genetics, Dec (online 29 Aug) 2012.
 Viruses and Other Gene Transfer Mechanisms is a related local webpage. Viruses and Other Gene Transfer Mechanisms is a related local webpage.
What'sNEW about HGT  | |
 The Tree of Life is a related local webpage. The Tree of Life is a related local webpage.
|
Membrane-bound organelles are a defining feature of eukaryotic cells, and play a central role in most of their fundamental processes. The Rab G proteins are the single largest family of proteins that participate in the traffic between organelles, with 66 Rabs encoded in the human genome. Rabs direct the organelle-specific recruitment of vesicle tethering factors, motor proteins, and regulators of membrane traffic. Each organelle or vesicle class is typically associated with one or more Rab, with the Rabs present in a particular cell reflecting that cell's complement of organelles and trafficking routes. |
A strikingly large repertoire of at least 20 Rabs appears to have been present in the last eukaryotic common ancestor (LECA), consistent with the 'complexity early' view of eukaryotic evolution. This comment comes from a collaboration of British and Swiss molecular biologists who classified from the wide eukaryotic kingdom certain proteins that participate in the traffic between organelles (see box). They reiterate, Examining the patterns of Rab evolution demonstrates the striking degree of Rab complexity in the last eukaryotic common ancestor (LECA).
While the "complexity early" hypothesis is gaining support, it has no ready explanation under strict darwinian theory. By darwinian logic, life should start simple and only later become complex. Meanwhile, "complexity early" is a basic requirement in cosmic ancestry.
 Tobias H Klöpper et al., "Untangling the evolution of Rab G proteins: implications of a comprehensive genomic analysis" [Open Access abstract], doi:10.1186/1741-7007-10-71, v10 n71, BMC Biology, 8 Aug 2012. Tobias H Klöpper et al., "Untangling the evolution of Rab G proteins: implications of a comprehensive genomic analysis" [Open Access abstract], doi:10.1186/1741-7007-10-71, v10 n71, BMC Biology, 8 Aug 2012.
 Metazoan Genes Older Than Metazoa? is a related local webpage. Metazoan Genes Older Than Metazoa? is a related local webpage.
 Genes Older Than Earth? is a related local webpage. Genes Older Than Earth? is a related local webpage.
Although our 20,000 genes occupy only 2% of the human genome, Almost 80% of the genome is biochemically active..., according to a longterm research project, the Encyclopedia of DNA Elements (ENCODE). In addition, large stretches of DNA that appeared to serve no functional purpose in fact contain about 400,000 regulators, known as enhancers, that help activate or silence genes, even though they sit far from the genes themselves. This news reinforces our view that genetic programming is complicated.
 The ENCODE Project Consortium, "An integrated encyclopedia of DNA elements in the human genome" [link], Nature, 6 Sep 2012. The ENCODE Project Consortium, "An integrated encyclopedia of DNA elements in the human genome" [link], Nature, 6 Sep 2012.
 Gautam Naik and Robert Lee Hotz, "'Junk DNA' Debunked" [link], pA3, The Wall Street Journal, 6 Sep 2012. Gautam Naik and Robert Lee Hotz, "'Junk DNA' Debunked" [link], pA3, The Wall Street Journal, 6 Sep 2012.
 Elizabeth Pennisi, "ENCODE Project Writes Eulogy for Junk DNA" [summary], p1159-1161 v337, Science, 7 Sep 2012. Elizabeth Pennisi, "ENCODE Project Writes Eulogy for Junk DNA" [summary], p1159-1161 v337, Science, 7 Sep 2012.
 Human Genome Is Much More Than Just Genes by Elizabeth Pennisi, ScienceNow, 5 Sep 2012. Human Genome Is Much More Than Just Genes by Elizabeth Pennisi, ScienceNow, 5 Sep 2012.
 ENCODE: The human encyclopaedia by Brendan Maher, Nature News, 5 Sep 2012. ENCODE: The human encyclopaedia by Brendan Maher, Nature News, 5 Sep 2012.
 Sequencing the Genome is a related section of our webpage "Can the Theory Be Tested?. Sequencing the Genome is a related section of our webpage "Can the Theory Be Tested?.
 Conserved Non-Genic Sequences is a related local webpage. Conserved Non-Genic Sequences is a related local webpage.
 Human Genome Search... is a loosely related local webpage. Human Genome Search... is a loosely related local webpage.
 09 Feb 2019: Rebuttal from Doolittle and Brunet. 09 Feb 2019: Rebuttal from Doolittle and Brunet.
 The Curiosity rover landed in Gale Crater on Mars, at 1:32AM EDT. NASA engineers hugged each other and cried.
The Curiosity rover landed in Gale Crater on Mars, at 1:32AM EDT. NASA engineers hugged each other and cried.
 Curiosity Rover Lands Safely on Mars, The New York Times, 6 Aug 2012. Curiosity Rover Lands Safely on Mars, The New York Times, 6 Aug 2012.
 Mars Landing: NASA's Curiosity Rover Safely Touches Down, ABC News, 6 Aug 2012. Mars Landing: NASA's Curiosity Rover Safely Touches Down, ABC News, 6 Aug 2012.
 Curiosity sets down safely on Mars by Eric Hand, Nature, 6 Aug 2012. Curiosity sets down safely on Mars by Eric Hand, Nature, 6 Aug 2012.
 Thanks, Chip Morrison, for midnight comments; and Pam Pacelli, for a facebook message. Thanks, Chip Morrison, for midnight comments; and Pam Pacelli, for a facebook message.
 Life on Mars! is a related local webpage. Life on Mars! is a related local webpage.
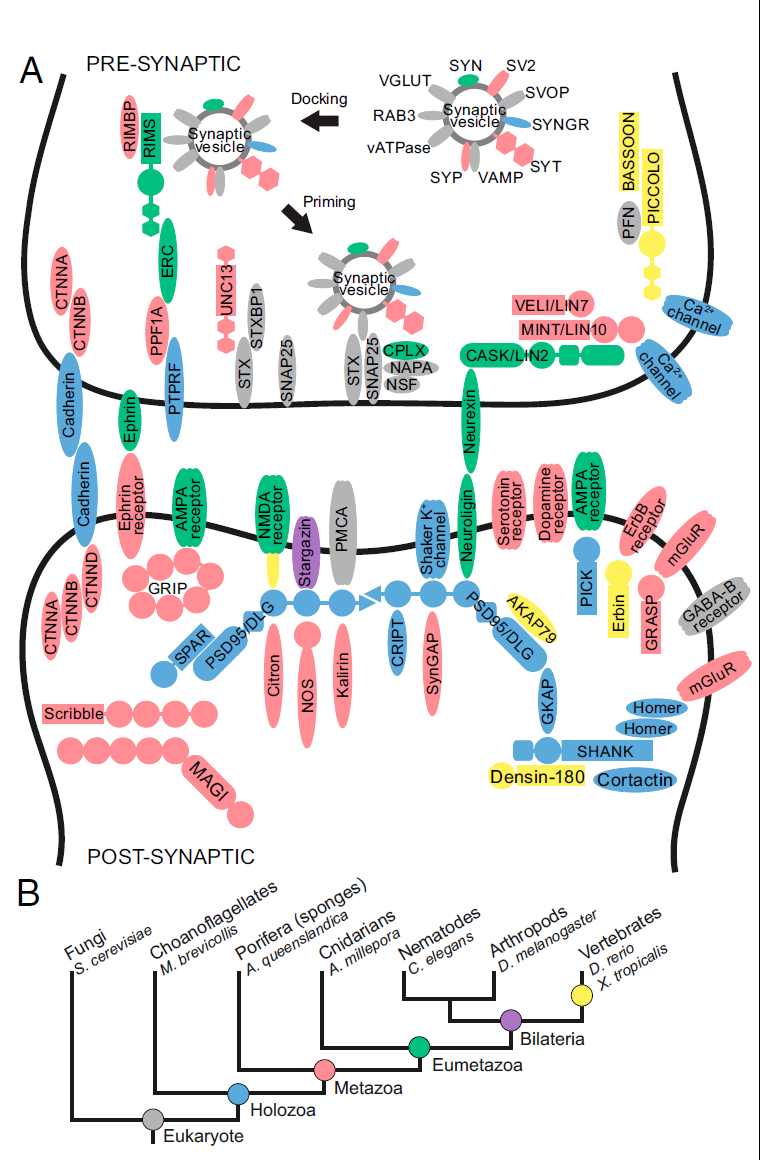
Surprisingly, the genome of the Poriferan demosponge, Amphimedon queenslandica, contains an almost complete set of genes homologous to those found in mammalian synapses..., although the organism does not assemble any structure morphologically resembling a synapse....
The accompanying illustration of a synapse includes the major genes that enable synapses to function, with their products in the roughly appropriate locations, if applicable. All of the pink ones represent genes with close homologs in sponges. But sponges have no synapses.
...We were hoping to understand why the marine sponge, despite having almost all the genes necessary to build a neuronal synapse, does not have any neurons at all, comments one of the lead authors of the new report. She should be wondering how the genes got there. In the darwinian scheme, they are entirely out of order. Genes are supposed to be composed incrementally, under selection pressure. But in sponges, with no nervous systems, what selection pressure could compose the genes for nervous systems?
In cosmic ancestry, genes precede the features they encode. Finding them in species that cannot yet deploy them is no surprise to us.
 Cecilia Conaco, Danielle S. Bassett et al., "Functionalization of a protosynaptic gene expression network" [abstract], doi:10.1073/pnas.1201890109, p10612-10618 v109 Suppl.1, Proc. Natl. Acad. Sci. USA, 26 Jun 2012. Cecilia Conaco, Danielle S. Bassett et al., "Functionalization of a protosynaptic gene expression network" [abstract], doi:10.1073/pnas.1201890109, p10612-10618 v109 Suppl.1, Proc. Natl. Acad. Sci. USA, 26 Jun 2012.
 Clues to Nervous System Evolution Found in Nerve-less Sponge, UC Santa Barbara (also EurekAlert!), 18 Jun 2012. Clues to Nervous System Evolution Found in Nerve-less Sponge, UC Santa Barbara (also EurekAlert!), 18 Jun 2012.
 Metazoan Genes Older Than Metazoa? is a related local webpage. Metazoan Genes Older Than Metazoa? is a related local webpage.
 Genes Older Than Earth? is a possibly related local webpage. Genes Older Than Earth? is a possibly related local webpage.
 Pedro Pereira replies and discussion ensues, 24-25 Jul 2012. Pedro Pereira replies and discussion ensues, 24-25 Jul 2012.
A lack of experimental data is no longer the primary limiting factor for researchers. Instead, it's how to make sense of what they already know. Stanford University's news office makes this comment following computer programming that models the complete life span of a free-living cell, Mycoplasma genitalium. This is a parasitic bacterium that can persist in human urogenital and respiratory tracts. Its genome is the smallest of any free-living organism — only 525 genes. (The common laboratory bacterium E. coli has eight times as many.)
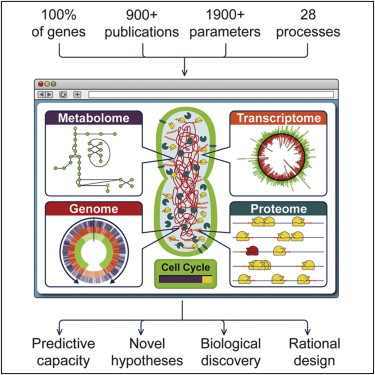 In spite of the genome's small size, running the model requires a cluster of 128 computers, and the simulation of a single cell division takes nine to ten hours. Interestingly, cell division for living Mycoplasma genitalium takes about as long.
In spite of the genome's small size, running the model requires a cluster of 128 computers, and the simulation of a single cell division takes nine to ten hours. Interestingly, cell division for living Mycoplasma genitalium takes about as long.
Comprehensive computer models of entire cells have the potential to advance our understanding of cellular function and, ultimately, to inform new approaches for the diagnosis and treatment of disease, comments James M. Anderson of the National Institutes of Health. We agree, but we notice no mention of possible contributions for evolution or origin-of-life research. Could those research programs be on the wrong track?
 Jonathan R. Karr, Jayodita C. Sanghvi et al., "A Whole-Cell Computational Model Predicts Phenotype from Genotype" [abstract], doi:10.1016/j.cell.2012.05.044, p389-401 v150, Cell, 20 Jul 2012. Jonathan R. Karr, Jayodita C. Sanghvi et al., "A Whole-Cell Computational Model Predicts Phenotype from Genotype" [abstract], doi:10.1016/j.cell.2012.05.044, p389-401 v150, Cell, 20 Jul 2012.
 Peter L. Freddolino and Saeed Tavazoie, "The Dawn of Virtual Cell Biology" [summary], doi:10.1016/j.cell.2012.07.001, p248250 v150, Cell, 20 Jul 2012. Peter L. Freddolino and Saeed Tavazoie, "The Dawn of Virtual Cell Biology" [summary], doi:10.1016/j.cell.2012.07.001, p248250 v150, Cell, 20 Jul 2012.
 Mark Isalan, "A cell in a computer" [abstract], doi:10.1038/488040a, p40-41 v488, Nature, 2 Aug 2012. Mark Isalan, "A cell in a computer" [abstract], doi:10.1038/488040a, p40-41 v488, Nature, 2 Aug 2012.
 Stanford researchers produce first complete computer model of an organism by Max McClure, Stanford University (+PhysOrg.com), 19 Jul 2012. Stanford researchers produce first complete computer model of an organism by Max McClure, Stanford University (+PhysOrg.com), 19 Jul 2012.
 In First, Software Emulates Lifespan of Entire Organism by John Markoff, The New York Times, 20 Jul 2012. In First, Software Emulates Lifespan of Entire Organism by John Markoff, The New York Times, 20 Jul 2012.
 Mark Isalan, "Systems biology: A cell in a computer" [html], doi:10.1038/488040a, p40-41 v488, Nature, 2 Aug 2012. Mark Isalan, "Systems biology: A cell in a computer" [html], doi:10.1038/488040a, p40-41 v488, Nature, 2 Aug 2012.
 Thanks for an alert, Stan Franklin. Thanks for an alert, Stan Franklin.
 Computer Models of Evolution and the four "Next" local webpages are related to this topic. Computer Models of Evolution and the four "Next" local webpages are related to this topic.
Arsenic-loving bacterium needs phosphorus after all
 Quirin Schiermeier, doi:10.1038/nature.2012.10971, [abstract], Nature News, 9 Jul 2012. Quirin Schiermeier, doi:10.1038/nature.2012.10971, [abstract], Nature News, 9 Jul 2012.
 Some bacteria can substitute arsenic for phosphorus, our notice of this report from a NASA-funded study, What'sNEW, 2 Dec 2010. Some bacteria can substitute arsenic for phosphorus, our notice of this report from a NASA-funded study, What'sNEW, 2 Dec 2010.
 Bacteria: The Space Colonists is a related local webpage. Search for "arsenic". Bacteria: The Space Colonists is a related local webpage. Search for "arsenic".
The evolution of RNA viruses is more promiscuous than we knew. A computational biologist and a botanical virologist make this observation in commentary on a study by a team of botanists, Liu et al. The studies begin by concentrating on dsRNA viruses, found in all domains of life but particularly characteristic in fungi. Ultimately the researchers observe rampant exchange of genes and gene fragments among all viruses and with their hosts. We note that in cosmic ancestry gene transfers supply the programs behind evolutionary progress. Koonin and Dolja write:
 ...The mosaic of these other key characteristics of viruses implies complex processes of multiple loss of ancestral genes (in particular, those for capsid proteins), gene exchange between distant viruses and transfer of viruses between distant hosts (such as fungi, plants, animals and protists, but possibly also between bacteria and fungi, even if only via endosymbiosis). ...The mosaic of these other key characteristics of viruses implies complex processes of multiple loss of ancestral genes (in particular, those for capsid proteins), gene exchange between distant viruses and transfer of viruses between distant hosts (such as fungi, plants, animals and protists, but possibly also between bacteria and fungi, even if only via endosymbiosis).
 A comprehensive history should integrate the histories of individual genes and can only adequately be conceptualized as a network of multiple evolutionary connections at the level of genes or even parts of genes encoding distinct protein domains. A comprehensive history should integrate the histories of individual genes and can only adequately be conceptualized as a network of multiple evolutionary connections at the level of genes or even parts of genes encoding distinct protein domains.
 Gene exchange that can lead to the emergence of new groups of viruses is not limited to viruses with the same type of genome structure. Gene exchange that can lead to the emergence of new groups of viruses is not limited to viruses with the same type of genome structure.
 ...Positive-strand RNA viruses might have pre-existed in archaeal ancestors of eukaryotes....If the archaeal host is confirmed, the impact of these findings on our understanding of RNA virus evolution will be dramatic. ...Positive-strand RNA viruses might have pre-existed in archaeal ancestors of eukaryotes....If the archaeal host is confirmed, the impact of these findings on our understanding of RNA virus evolution will be dramatic.
|
 Eugene V Koonin and Valerian V Dolja, "Expanding networks of RNA virus evolution" [abstract | html], doi:10.1186/1741-7007-10-54, n54 V10, BMC Biology, online 20 Jun 2012. Eugene V Koonin and Valerian V Dolja, "Expanding networks of RNA virus evolution" [abstract | html], doi:10.1186/1741-7007-10-54, n54 V10, BMC Biology, online 20 Jun 2012.
 Huiquan Liu et al., "Evolutionary genomics of mycovirus-related dsRNA viruses reveals cross-family horizontal gene transfer and evolution of diverse viral lineages" [Open Access abstract], doi:10.1186/1471-2148-12-91, v12 n91, BMC Evolutionary Biology, 20 Jun 2012. Huiquan Liu et al., "Evolutionary genomics of mycovirus-related dsRNA viruses reveals cross-family horizontal gene transfer and evolution of diverse viral lineages" [Open Access abstract], doi:10.1186/1471-2148-12-91, v12 n91, BMC Evolutionary Biology, 20 Jun 2012.
 Viruses and Other Gene Transfer Mechanisms is a related local webpage. Viruses and Other Gene Transfer Mechanisms is a related local webpage.
What'sNEW about HGT  | |
|


 In spite of the genome's small size, running the model requires a cluster of 128 computers, and the simulation of a single cell division takes nine to ten hours. Interestingly, cell division for living Mycoplasma genitalium takes about as long.
In spite of the genome's small size, running the model requires a cluster of 128 computers, and the simulation of a single cell division takes nine to ten hours. Interestingly, cell division for living Mycoplasma genitalium takes about as long.